All published articles of this journal are available on ScienceDirect.
The Prevalence, Molecular Characterization and Antimicrobial Susceptibility of S. aureus Isolated from Impetigo Cases in Duhok, Iraq
Abstract
Background:
Staphylococcus aureus is one of the most important opportunistic pathogens. Impetigo is the common contagious bacterial infection of the skin caused by S. aureus.
Method:
Samples were taken from 204 patients with impetigo disease. S. aureus isolates were tested for their antimicrobial susceptibility. Genomic DNA of S. aureus was used to transform E. coli HB101 strain and expression capability of S. aureus plasmids in transformed E. coli was investigated. 68.62% (140/204) of the specimens were nonbullous impetigo and 31.38% (64/204) were bullous impetigo. S. aureus strains were isolated from 41.66% (85/204) of impetigo cases (82.35% from nonbullous and 17.65% from the bullous impetigo). There was an inverse relationship between the incidence of S. aureus isolated and age.
Result:
Three biotypes of S. aureus were identified based on their fermentation of different sugars. All isolates were resistant to penicillin and most isolates were resistant to ampicillin (95.3%), amoxicillin, (94.11%) and cephalexin (90.95%). Most isolates were sensitive against vancomycin and rifampicin (98.83%). 5.88% (5/85) of S. aureus isolates were identified as MRSA. A maximum of 5 markers from S. aureus isolates were capable to be expressed in transformed E. coli HB101 strains. The incidence of impetigo caused by S. aureus is comparable with reports from elsewhere. S. aureus isolates showed multidrug resistance against antibiotics.
Conclusion:
Plasmids of S. aureus are capable to show its expression in E. coli HB101. Molecular study is needed to investigate the role of plasmids in different patterns of multi drug resistance.
INTRODUCTION
Staphylococcus aureus is a major pathogen responsible for both nosocomial and community acquired infections [1]. The severity of these infections varied from local simple wounds to severe systemic diseases. This pathogen causes a range of diseases, including skin abscesses, wound infections, osteomyelitis, endocarditis, pneumonia, bacteremia and toxemic syndromes [2]. Impetigo is a common contagious bacterial infection of the skin caused by S. aureus. This infection is mostly seen in children, especially in tropical countries [3]. Impetigo spreads from person to person through direct contact with the discharge from the sores [4]. Two types of impetigo are diagnosed, nonbullous impetigo which represents a host response to the infection, and it is characterized by reddened sores with honey-yellow crusting on them, and bullous impetigo which is caused by staphylococcal toxin and no host response is required to manifest clinical illness, and it is characterized by bullae large thin-walled blisters that contain clear or cloudy yellow fluid [4].
S. aureus is one of the most successful pathogens to acquire antibiotic-resistance mechanisms [5]. Methicillin-resistant S. aureus (MRSA) is an important pathogen causing poisoning and toxin mediated infections. The resistance in MRSA is related to a chromosomal mecA gene that specifies the production of an abnormal penicillin–binding protein called PBP2a, resulting in resistance not only to methicillin but to all β–lactam [6]. Most plasmids have a narrow host range but some can replicate in a wider host range [7]. The genetic exchange of plasmids containing antibiotic resistant genes among organisms is believed to play a crucial role in the evolution of antibiotic resistant bacteria. Staphylococcal β-lactamase is carried on plasmids and can be inducible or non-inducible with antibiotic contact [8]. Plasmid mediated drug resistance in S. aureus to chloramphenicol, gentamycin, tobramycin and kanamycin has been documented [9]. The aim of this study was to determine the occurrence of impetigo cases and the occurrence of multidrug resistant S. aureus in these cases in Duhok city. Additionally, this work aimed to examine the expression ability of S. aureus plasmids in E. coli HB101 by transformation.
MATERIALS AND METHOD
Sample Collection
A total of 204 specimens were collected from patients with impetigo disease at Azadi General Hospital in Duhok city from November 2008 to April 2009. Samples were collected either by using sterile disposable cotton swabs moistened in brain heart infusion (BHI) broth or by sterile forceps from infected skin. The study was conducted with the approval of ethics committee of the University of Duhok, School of Science.
Identification and Characterization
The samples were cultured in BHI broth and incubated overnight at 37˚C. Samples were subcultured on blood agar and mannitol salt agar (MSA) and incubated at 37˚C for overnight. The isolates were identified as S. aureus based on morphology, Gram stain, catalase test, coagulase test, DNase test and mannitol salt agar fermentation. Suspected S. aureus colonies were further identified using API® Staph Kit (Biomerieux, France).
Antibiotic Sensitivity Test
All S. aureus isolates were tested against 13 different antimicrobial drugs (Bioanalyse, Turkey) as listed in Table 1. Screening for antibiotic resistance was performed using disk diffusion assay according to Bauer et al. [10]. Bacterial suspensions were prepared in 1.0 ml of sterile saline solution. 0.5 ml of suspensions (optical density of 0.3 at 600nm) was spread on Mueller-Hinton agar (Oxoid Limited, Hampshire, England) and then incubated at 37˚C for 24 hours. The inhibition zones were measured according to the recommendations of the National Committee for Clinical Laboratory Standards (NCCLS) guidelines [11].
| Class | Antimicrobial | Symbol | Disk potency/ µg | |
|---|---|---|---|---|
| β-lactams | Penicillins | Penicillin | PG | 10 |
| Amoxicillin | Ax | 30 | ||
| Methicillin | Met | 5 | ||
| Ampicillin | AMP | 30 | ||
| Cephalosporins | Cephalexin | CL | 30 | |
| Cephotaxime | CTX | 30 | ||
| Fluoroquinolones | Ciprofloxacin | CIP | 5 | |
| Macrolides | Erythromycin | E | 15 | |
| Fusidanes | Fusidic Acid | FA | 10 | |
| Lincosamides | Lincomycin | L | 2 | |
| Rifamycin | Rifampicin | RA | 5 | |
| Sulfonamides | Trimethoprim | TMP | 5 | |
| Glycopeptides | Vancomycin | VA | 30 | |
Plasmid Extraction and Gel Electrophoresis
Plasmid DNA of S. aureus was isolated according to Lema et al. [12]. Briefly, S. aureus cells were harvested from an overnight 10 ml culture and the pellet was suspended in 1ml TE buffer pH 7.4 (10mM of Tris-base and 1mM of EDTA). Suspended cells were distributed into 1ml tubes and centrifuged at 6000 rpm for 5 minutes. The pellet was suspended in 200μl acetone and incubated on ice for 5 minutes then centrifuged at 6000 rpm for 5 minutes. The pellet was resuspended in 100μl of TE buffer (contain lysozme 2mg/ml) and incubated at 37˚C for 1 h. 300μl of TESN (lysis) buffer with pH 7.4 (10mM of Tris-base, 1mM of EDTA, 1% of SDS and 6 M of NaI) as added and incubated at 65˚C for 5 minutes, then 300µl isopropanol was added and incubated for 5 minutes at room temperature. The mixture was centrifuged at 6000 rpm for 2 minutes and the pellet was resuspended in 500 μl of isopropanol 40% and centrifuged at 10000 rpm for 10 minutes. The pellet was resuspended in 1ml of ethanol 70% and centrifuged at 10000 rpm for 10 minutes. Finally, the pellet was suspended in 100μl of TE buffer and stored at -20˚C. The DNA was analyzed by 1.5% agarose gel electrophoresis (80 V in 1× TAE buffer).
Transformation
DNA transformation was achieved according to Sambrook et al. [13]. The preparation of the competent cells and DNA uptake were achieved as follows:
Preparation of Competent Cells
Single colony of Escherichia coli HB101 strain was inoculated into 100ml nutrient broth and incubated at 37˚C for 3h and then was incubated on ice for 10 minutes. Cells were harvested by centrifuge at 4000 rpm for 10 minutes at 4˚C. The pellet was suspended in 10 ml of 0.1M calcium chloride. The mixture was centrifuged at 4000 rpm for 10 minutes at 4˚C. The pellet was suspended in 2 ml of 0.1M calcium chloride and kept on ice.
DNA Uptake
10μl of the genomic DNA of S. aureus was mixed with 100μl of the competent cells of E. coli and incubated on ice for 30 min. Tubes were exposed to heat shock at 45˚C for 90 seconds, then to ice for 1-2 minutes. 800μl of nutrient broth was added and incubated at 37˚C for 45 minutes. Transformed E. coli colonies were screened on nutrient agar containing 100mg/ml penicillin. Transformed cells were tested for their acquisition of resistance antibiotics. The susceptibility profile of transformants was compared with that of the S. aureus isolate and E. coli HB101 strain used in transformation.
RESULTS
Isolation and Characterization of S. aureus
A total of 204 specimens were collected from patients clinically diagnosed with impetigo at Azadi general hospital in Duhok city. Out of 204 cases, 140 (68.62%) specimens were nonbullous impetigo and 64 (31.38%) were bullous impetigo. Prevalence rate of S. aureus was 41.66% (85/204). The incidence of S. aureus isolated from nonbullous impetigo included 70 isolates (82.35%) and 15 isolates (17.65%) were from the bullous impetigo specimens. Twenty isolates were selected randomly and confirmed as S. aureus by API Staph System test. Based on their fermentation to different sugars, these twenty isolates were classified into three biotypes (Table 2).
| No. of isolate (%) | GLU | FRU | MNE | MAL | LAC | TRE | MAN | XLT | MEL | NIT | PAL | VP | RAF | XYL | SAC | MDG | NAG | ADH | URE |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 11 (55) | + | + | + | + | + | + | + | - | - | + | + | + | - | - | + | + | - | - | + |
| 6 (30) | + | + | + | + | + | + | + | - | - | - | + | + | + | - | + | + | - | + | - |
| 3 (15) | + | + | + | + | + | + | + | - | - | + | - | - | + | - | + | + | + | - | - |
The percentage of S. aureus prevalence in age group from few months to 5 years was 50.66% and increased to 55.55% in the age group of 6-10 years. Then the percentages were continually decreased to 14.3% in the age group of more than 30 years (Table 3).
| Age group years | No. of impetigo cases | No. of S. aureus isolates (%) |
|---|---|---|
| 0-5 | 75 | 38 (50.66) |
| 6-10 | 54 | 30 (55.55) |
| 11-20 | 43 | 12 (27.9) |
| 21-30 | 25 | 4 (16) |
| ≥ 31 | 7 | 1 (14.3) |
Antibiotic Susceptibility Test
All 85 isolates of S. aureus were tested for their susceptibility against thirteen different antibiotics. Results indicated that all isolates were resistant to penicillin G, and most isolates were resistant to ampicillin (95.3%), amoxicillin (94.11%) and cephalexin (90.95%). On the other hand, most isolates were sensitive against vancomycin and rifampicin (98.83%) (Table 4).
| Antibiotics | No. of resistant isolates (%) |
|---|---|
| Penicillin | 85 (100) |
| Ampicillin | 81 (95.3) |
| Amoxicillin | 80 (94.11) |
| Cephalexin | 77 (90.58) |
| Lincomycin | 57 (67.05) |
| Cefotaxime | 41 (48.23) |
| Erythromycin | 10 (11.76) |
| Ciprofloxacin | 5 (5.88) |
| Methicillin | 5 (5.88) |
| Trimethoprim | 3 (3.52) |
| Fusidic Acid | 2 (2.35) |
| Vancomycin | 1 (1.17) |
| Rifampicin | 1 (1.17) |
Out of 85 isolates, 5(5.88%) were found to be methicillin resistant S. aureus (MRSA).
The Plasmid Profile of S. aureus Isolates
In our study, 10 different isolates showed multi drug resistance (MDR) against at least three different antibiotics. These isolates were selected and the plasmid profile was investigated. All tested isolates contained two small plasmids represented by two bands (Fig. 1).
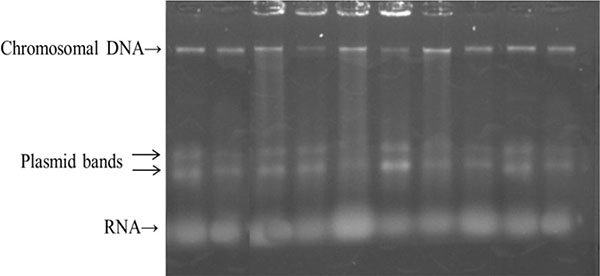
Transformation
In order to investigate the expression ability (antibiotic resistance) of the genetic marker in E. coli, genomic DNA from MDR S. aureus isolates was used to transform E. coli HB101 rifampcin resistant competent cells. All ten antibiotic resistant markers were checked for their transformation to E. coli HB101. Transformants E. coli were screened on nutrient agar supplemented with different antibiotics. Results showed that only a maximum of 5 markers (penicillin, ampicillin, amoxicillin, cephalexin and lincomycin) were capable to transfer transformation from S. aureus isolates which have different patterns of MDR (Table 5).
| Antibiotic resistant | S. aureus | E. coli HB101 | E. coli HB101Transformation |
|---|---|---|---|
| Penicillin resistant | ve+ | ve- | ve+ |
| Ampicillin resistant | ve+ | ve- | ve+ |
| Amoxicillin resistant | ve+ | ve- | ve+ |
| cephalexin resistant | ve+ | ve- | ve+ |
| Lincomycin resistant | ve+ | ve- | ve+ |
| Rifampcin resistant | ve- | ve+ | ve+ |
DISCUSSION
Although S. aureus is an important pathogen, many healthy people carry it as part of the nose, throat, perineum and skin microbiota [14]. Our results were in agreement with those obtained from other studies which showed that non-bullous impetigo was the most common form of impetigo and that S. aureus was the main bacterium that causes non-bullous impetigo [4, 15]. In studies conducted in Kurdistan region, S. aureus isolated from different clinical and environmental sources were also characterized into three biotypes [16, 17]. Results revealed that there is an inverse relationship between the percentage of isolated S. aureus and patient age. Similar patterns of these results were observed in different studies conducted in Iraq and worldwide [18, 19].
Overuse and misuse of antibiotics have led to increased levels of antibiotics resistance [20-22]. Antibiotic susceptibility of isolated S. aureus was found to be comparable with other studies conducted locally and worldwide in which S. aureus clinical isolates from different hospitals and medical institutions were found to be highly resistant to penicillin G (92% to 100%) and highly sensitive to vancomycin [16, 17, 23]. High resistance to penicillin is due to the ability of S. aureus to produce β- lactamase enzymes [24]. It is known that most S. aureus clinical isolates are β-lactamase producers [23]. S. aureus is also highly sensitive against rifampcin because this antibiotic specifically inhibits bacterial RNA polymerase [25].
MRSA is an increasing problem worldwide in community and hospital environment [26-29]. MRSA colonization rate varied in different countries from about 1% to 35% [22, 26, 27, 30-34]. Combination of host, geographical, environmental and bacteria factors could contribute to the nasal carriage of S. aureus and MRSA among population. There is evidence suggesting that development of antibiotic resistant human microflora in hospitals is mainly caused by the use of antimicrobial agents by the medical profession [35]. The health care workers usually act as asymptomatic carriers of multiple drug resistance organisms, especially MRSA, and help in its transmission to patients [36].
Studies with S. aureus clinical isolates showed that these isolates harbored many plasmids of different sizes. Kloose et al. [37] described plasmids with sizes between 2-29kb. Another plasmid profile study was conducted with 88 strains of S. aureus isolated causing impetigo cases and showed that most strains harbor different size plasmids between 1.29-25 kb [38].
S. aureus plasmids were found to be expressed in E. coli HB101. These results were in agreement with those obtained from different studies carried out worldwide [23, 39, 40]. A study showed that 23 kb plasmid from a selected multidrug resistant strain of S. aureus was used to transform a sensitive E. coli LE 392 and it was found that E. coli LE 392 which was sensitive to ampicillin, amoxycillin and penicillin before transformation became resistant to these drugs [39]. Results revealed that transformants E. coli acquired resistance against penicillin, ampicillin, amoxicillin, cephalexin and lincomycin. Resistance to penicillin, ampicillin and amoxicillin is mediated by beta-lactamases, which may also act against first generation cephalosporins, such as cephalexin. Thus, a single genetic marker transferred from S. aureus to E. coli could be responsible for the acquisition of resistance to these 4 antibiotics. It is revealed that five S. aureus plasmids, coding for tetracycline or chloramphenicol resistance, have been transformed, replicated and expressed into Bacillus subtilis [41]. It is believed that the multiple drug resistance of S. aureus is due to several drug resistance genes in a single plasmid each with its own different resistance markers [42].
It is worth mentioning that other mobile genetic elements may have been incorporated to E. coli cell and even into its chromosome, and more in-depth studies would be needed to identify this event. Furthermore, it would be important to detect the presence of the plasmid in the transformed E. coli cell, confirming that the acquired resistance is actually related to the presence of the plasmid. There are many factors that play role in selection of the multi-drug resistant strains such as drug misuse, prescription of antibiotic without culturing, identification, and determination of susceptibility tests of pathogen, and giving of broad–spectrum antibiotics instead of narrow-spectrum antibiotics.
CONCLUSION
This study concluded that the incidence of impetigo caused by S. aureus is comparable with reports from elsewhere. Most S. aureus isolates showed multidrug resistant phenomena against different antibiotics. Plasmids from these multidrug resistant isolates were capable to transform E. coli HB101 strain into multidrug resistant bacteria. Further molecular studies are needed to investigate the role of plasmids in different patterns of MDR.
CONFLICT OF INTEREST
The authors confirm that this article content has no conflict of interest.
ACKNOWLEDGEMENTS
Declared none.


